Qing Wang, Min Pan, Jie Wei, Xiaoqing Liu, and Fuan Wang*
Key Laboratory of Analytical Chemistry for Biology and Medicine (Ministry of Education), College of Chemistry and Molecular Sciences, Wuhan University, Wuhan, 430072, P. R. China
ACS Sens., 2017, 2 (7), pp 932–939
DOI: 10.1021/acssensors.7b00168
Publication Date (Web): June 14, 2017
Abstract:
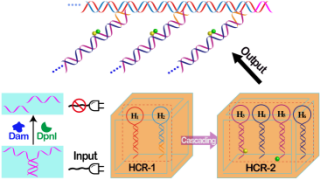
Methyltransferase (MTase)-catalyzed DNA methylation plays a vital role in the biological epigenetic processes of key diseases and has attracted increasing attention, making the amplified detection of MTase activity of great significance in clinical disease diagnosis and treatment. Herein, we developed an isothermal, enzyme-free, and autonomous strategy for analyzing MTase activity based on concatenated hybridization chain reaction (C-HCR)-mediated Förster resonance energy transfer (FRET). In a typical C-HCR procedure without MTase (Dam), Y-shaped initiator DNA activates upstream HCR-1 to assemble a double-stranded DNA (dsDNA) copolymeric nanowire consisting of multiple tandem DNA trigger units that motivate downstream HCR-2 to successively bring a fluorophore donor/acceptor (FAM/TAMRA) pair into close proximity, leading to the generation of an amplified FRET readout signal. The target Dam MTase and auxiliary DpnI endonuclease can sequentially and specifically recognize/methylate and cleave the Y-shaped initiator oligonucleotide, respectively, and thus prohibit the C-HCR process and FRET signal generation, resulting in the construction of a signal-on sensing platform for MTase assay. Our proposed isothermal enzyme-free C-HCR amplification approach was further utilized for screening MTase inhibitors. Furthermore, the proposed C-HCR approach can be easily adapted for probing other different MTases and for screening the corresponding inhibitors just by changing the recognition sequence of Y-shaped initiator DNA through a “plug-and-play” format. It provides a versatile and robust tool for highly sensitive detection of various biotransformations and thus holds great promise in clinical assessment and diagnosis.
Full text: http://pubs.acs.org/doi/abs/10.1021/acssensors.7b00168
