Professor Chen Suming from Wuhan University's Institute for Advanced Studies and his team have made a significant breakthrough in precise mass spectrometry analysis of isomers, with their latest research, XL-MSDigger: a deep learning-based, versatile solution for cross-linking mass spectrometry, published in Nature Communications.
The team was the first to apply deep learning based on the self-attention mechanism to predict multidimensional peptide information, developing the Deep4D model using a Transformer architecture.
This model enabled the joint prediction of collision cross-section (CCS), retention time, and fragment intensity for various modification types, such as phosphorylated peptides, achieving a median relative error of only 0.72 percent in CCS prediction and reducing the false-positive rate in modification site isomer identification (Anal. Chem. 2023, 95, 7495-7502).
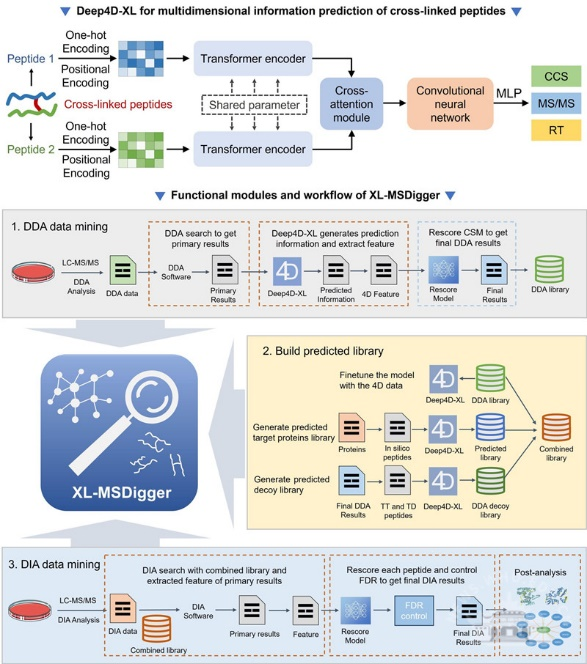
The team further developed the Deep4D-XL deep learning model (Nat. Commun. 2026, 17, 2554), introducing a twin-network structure to characterize the two peptide segments of cross-linked peptides, integrating features through a cross-attention module, and finally outputting multidimensional predictions via a decoding module.
The model achieved a retention time prediction correlation of R²= 0.97 with experimental values, a median relative error of 1.52 percent in CCS prediction, and a median point product of 0.88 for fragment ion intensity prediction.
Building on Deep4D-XL, the team developed the XL-MSDigger platform, which supports both data-dependent acquisition (DDA) and data-independent acquisition (DIA) modes for intelligent cross-linking mass spectrometry analysis.
In DDA analysis, the platform extracts 16 features using multidimensional predictive information and applies deep neural network rescoring, resulting in a 107 percent increase in cross-linked peptide spectrum matches and a 76.3 percent increase in protein-protein interaction identifications.
The team also established the first systematic framework for evaluating cross-linked peptide false discovery rates, achieving a fivefold increase in identifications through rescoring and identifying 90 protein-internal cross-linked peptides and several high-confidence protein-protein interactions not detected in DDA analysis.